March 17, 2026

The traditional path from target identification to preclinical candidate nomination averages 4.5 years, in part because medicinal chemistry is constrained by how many molecules can be designed, synthesized, and tested in iterative cycles. AI-driven platforms are slashing that to 12–18 months. The key lies in generative chemistry: machine learning models that design, evaluate, and optimize novel molecules computationally before a chemistry team commits to extensive synthesis in a lab.
Insilico Medicine illustrates the shift from hypothesis generation to candidate nomination at scale. Using its Pharma.AI platform, the company advanced 20 preclinical candidates between 2021 and 2024, synthesizing only 60–200 molecules per program. Its lead compound, Rentosertib, became the first fully AI-designed drug to reach Phase II clinical trials for pulmonary fibrosis, earning FDA Orphan Drug designation [2].
Exscientia (now merged with Recursion in a $688M deal) set a different kind of milestone with DSP-1181, an OCD candidate widely cited as the first AI-designed drug to enter a Phase I clinical trial, moving from concept to trial start in just 12 months. Even though that product did not proceed through later phases, it established a landmark proof of concept: AI-driven design can compress the early drug-discovery cycle without sacrificing the rigor required to reach first-in-human testing, and it helped catalyze the next wave of AI-enabled drug development.
The shift is now showing up as productized platforms. Companies like Numerion Labs, and Axiom Vault are offering systems that can screen millions of molecules against a set of defined constraints in minutes, turning what used to be a bottleneck into something closer to an interactive step in the workflow. If the first wave of AI value was speed in target identification, the next question is reliability: how consistently can these faster “first-in-human” bets translate into clinical efficacy?
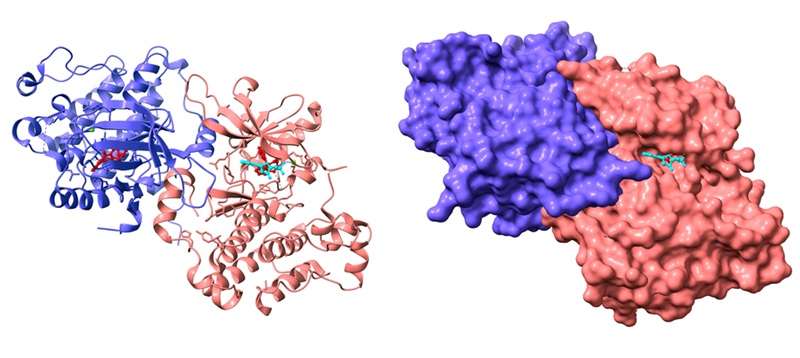
Perhaps the most transformative AI contribution came from protein structure prediction. DeepMind and EMBLE-EBI’s AlphaFold predicted the 3D structures of over 200 million proteins and made them freely available. Protein structure is a cornerstone of drug discovery: if you can see the shape of a target, you can design (or select) molecules that are more likely to bind the right place. Recognizing the impact of this breakthrough, David Baker, Demis Hassabis, and John Jumper, the leaders behind AlphaFold’s advance in protein structure prediction and computational protein design, received major scientific honors, including the 2024 Nobel Prize in Chemistry, as well as the 2023 Lasker Basic Medical Research award and the 2023 Breakthrough Prize in Life Sciences for their work. The more recent AlphaFold3 model has been updated specifically for drug discovery. It shines in areas where a usable structure did not previously exist for a target, enabling faster structure-based hypothesis generation and screening.
AlphaFold’s open approach has lowered barriers for academic groups and smaller biotech companies, creating shared infrastructure that is accelerating early discovery across the ecosystem. As of the writing of this article, a pubmed search for AlphaFold returned almost 3,000 publications.
In contrast to open models, proprietary systems are being positioned as end-to-end “drug discovery engines” that can be licensed to large pharmaceutical partners. Google’s spinoff, Isomorphic Labs, represents this move towards commercialization and exclusivity. Leveraging IsoDDe, a proprietary ‘drug discovery engine’, they expect to have its first oncology drug in clinical trials this year.
At the same time, open‑source approaches are still being actively developed, with researchers at MIT creating Boltz-2, extending modeling capability and binding-affinity prediction with improved computational efficiency, another sign that the field is advancing on both, the open and closed fronts at once . With speed of discovery well established, the real test will be whether these tools can deliver safer, faster, and more successful drugs that actually make it into the clinic.

Early analyses suggest that AI-discovered compounds show 80–90% success rates in Phase I trials, compared to 40–65% for conventional development, with cost savings of up to 70% [3]. A major application area of these tools is drug repurposing.
During the COVID-19 pandemic, BenevolentAI’s knowledge graph rapidly identified an existing drug approved for rheumatoid arthritis, baricitinib, as a potential candidate for repurposing to treat COVID, which subsequently received emergency authorization. By early 2026, AI-drug-discovery companies had expanded collaborations across multiple therapeutic areas, aiming to compress the distance from bench to bedside. This is where additional stakeholders matter: the story isn’t just “AI company + biopharma,” it’s also regulators, trial operators, data stewards, and the clinical sites that must execute (and trust) the work. Regulators are now active participants in shaping what “credible AI” looks like in drug development, as FDA and EMA have articulated principles for the use of AI across the medicine lifecycle. While Health Canada has published guidance on the use of AI in medical devices, similar guidance on the use of AI in drug development has not been formally established.

Most AI-discovered compounds remain in early-stage trials; very few have completed the full journey to approved medicines. AI excels at target identification and molecule optimization, but late-stage clinical trials, which involve testing safety and efficacy in large human populations, remain expensive, slow, and frequently disappointing regardless of how a compound was discovered [3].
Data quality is a persistent bottleneck: models are constrained by incomplete datasets, measurement noise, and biases toward well-studied diseases. AI can propose candidates that look promising computationally but fail in biological reality, highlighting the persistent gap between in silico prediction and in vivo validation.
And beyond the science, there’s a governance challenge: documentation, validation, and monitoring expectations are still maturing. For many organizations, the hard part won’t be buying and incorporating a model, it will be building a defensible process around it.
AI is not replacing medicinal chemists, it is giving them radically better starting points and shrinking the time spent in unproductive synthesis. But moving from molecule to medicine still depends on the same non-negotiables: reproducible biology, efficacy, rigorous clinical trials, manufacturable programs, and getting regulators to trust the evidence.
In the next few years, the winners won’t be the groups with the flashiest models; they’ll be the ones who connect prediction to proof end-to-end, and do it transparently enough that regulators, clinicians, and patients can believe the results. The molecule may “design itself” sooner than we think but getting it safely to the bedside will remain a team sport.
Disclaimer: The mention of specific companies, products, or organizations in this article is for informational purposes only and does not imply endorsement. The companies whose products were referenced were not consulted, involved in the preparation of this content, nor did they provide any funding or compensation.
References
[2] Z. Xu et al., “A generative AI-discovered TNIK inhibitor for idiopathic pulmonary fibrosis: a randomized phase 2a trial.,” Nat. Med., vol. 31, no. 8, pp. 2602–2610, Aug. 2025, doi: 10.1038/s41591-025-03743-2.
[3] M. Kp Jayatunga, M. Ayers, L. Bruens, D. Jayanth, and C. Meier, “How successful are AI-discovered drugs in clinical trials? A first analysis and emerging lessons.,” Drug Discov. Today, vol. 29, no. 6, p. 104009, Jun. 2024, doi: 10.1016/j.drudis.2024.104009.